|
Remaining errors were corrected by visual inspection of a mapping of all reads against the Medaka-polished consensus using CLC Genomics Workbench v22.0.1. From this read mapping, a consensus sequence with an average coverage of 116x (range 24x to 192x) was extracted, which was polished using Medaka v1.4.3. The final libraries were eluted in 15 μL Elution Buffer (EB) and 70-120 ng of each library was loaded onto an R9.4.1 flow cell, which were run for 72 hours.ĭata from all three runs was mapped to Monkeypox virus strain Israel_2018 (GB: MN648051.1) using CLC Genomics Workbench v22.0.1. During this clean-up, two of the DNA samples were washed with Long Fragment buffer (LFB), while one sample was washed with Short Fragment Buffer (SFB). AMX adapter was ligated to the end-prepped DNA as per the SQK-LSK109 kit protocol, followed by a 0.4x AMPure bead clean-up. The end-prepped products were purified using a 1x AMPure bead clean-up with two 80% EtOH washes. We started with 400 ng of each DNA extract in 25 μL, which was end-prepped using the NEBNext Ultra II End Repair/dA-tailing module kit. Sequencing libraries were prepared using the ligation sequencing kit (SQK-LSK109) from Oxford Nanopore Technologies (ONT). Subsequently, a conventional PCR was performed with Monkeypox-specific primers, confirming the presence of Monkeypox virus DNA in each extract. Viral DNA was extracted from each swab sample (three in total) using the QIAamp DNA Mini Kit (QIAGEN) according to the manufacturer’s instructions. To access these videos, click here.Bert Vanmechelen, Tony Wawina-Bokalanga and Piet Maes.Ī Monkeypox virus genome was obtained from a second Belgian case at the Rega Institute, KU Leuven. Here, CDS annotations are visible (in yellow), one of which has been selected. On the right is a region of the same alignment after zooming in. The alignment on the left is the large scale view. Finally, Qiagen hosts webinars addressing the use of and updates to this software. Figure 1 Visualizing a whole genome alignment using the CLC Workbench viewer. Additionally, there are tutorials available for different workflows. For the CLC Genomics Workbench manual, click here. Getting Helpĭocumentation for the CLC Genomics Workbench is available under the Help tab in the software. Working with CLC Genomics Workbench requires login to the NIH network or VPN connection if remote. This software requires access to a floating license server (three simultaneous users), and so care should be taken to return the license when the software is not actively being used (i.e. You must submit a request through to obtain access to CLC Genomics Workbench. Visualizations can be exported with varying resolutions as the following:. 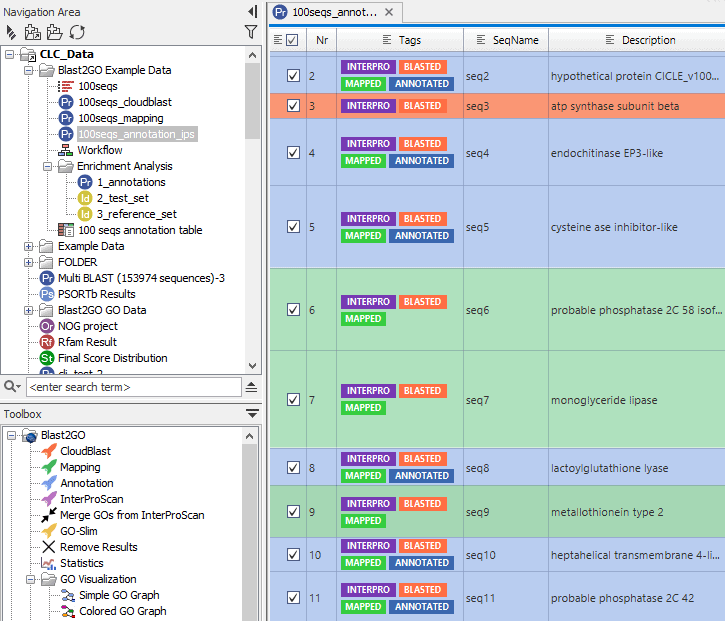
We can import data to CLC Genomics Workbench from the following sequencing instruments: 
Tracks/annotations from the UCSC Genome Browser and COSMIC database.Things to KnowĪlthough CLC Genomics Workbench is comprehensive, compute resources on a user's local machine may be a limiting factor for analysis. The combination of CLC Genomics Workbench and IPA allows us to go from a gene level of understanding to understanding biological function and regulatory mechanisms. Results from CLC Genomics can be imported to IPA for pathways analysis. Obtain protein structure from PDB and sequences from UniProtĬLC Genomics Workbench and Ingenuity Pathway Analysis (IPA) are both products developed by Qiagen. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
